Part:BBa_K510000:Experience
In order to characterize the function of the miniTn7BB-Gm minitransposon, we performed a set of experiments aimed to (i) determine whether miniTn7BB-Gm can transpose efficientely in an enterobacterial host (E. coli DH5α) and a non-enterobacterial host (P. putida KT2440), (ii) measure the transposition frequency when miniTn7BB-Gm was delivered by electroporation, chemical transformation and mating, (iii) determine whether insertion occurs site-specifically at the chromosomal attTn7 site, and (iv) determine whether flipase expression in trans can excise the drug resistance marker in the chromosomal copy of miniTn7BB-Gm.
Contents
Transposition of miniTn7BB-Gm in P. putida
In order to demonstrate miniTn7BB-Gm transposition into the P. putida KT2440 chromosome, pUC18Sfi-miniTn7BB-Gm was transferred to this host by two different means: (i) co-electroporation with the helper plasmid pTNS2 [pTNS2 is not replicative in P. putida, but transiently produces the Tn7 transposition machinery ([http://http://www.nature.com/nmeth/journal/v2/n6/abs/nmeth765.html Choi et al., 2005]), and (ii) tetraparental mating, using DH5α/pUC18Sfi-miniTn7BB-Gm as the donor, DH5α/pRK2013 as helper to provide the Tra functions, DH5α/pTNS2 to provide the Tn7 transposase and P. putida KT2440 as the recipient. P. putida clones bearing miniTn7BB-Gm insertions were selected on LB plates supplemented with Gm (and chloramphenicol when mating was used). Viable counts were also performed in both experiments, and plasmid transformation efficiency was determined using the gentamycin resistant replicative plasmid pBBR1mcs-5 in the electroporation experiment. The results are shown in Table 1.
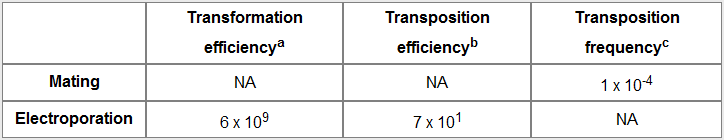
Table 1. Characterization of miniTn7BB-Gm transposition in P. putida KT2440. (a) Transformant (gentamycin resistant) cfu / μg pBBR1-mcs5. (b) cfu bearing a miniTn7BB-Gm insertion (gentamycin resistant) / μg pUC18Sfi-miniTn7BB-Gm. (c) Recipient cfu bearing a miniTn7BB-Gm insertion (gentamycin and chloramphenicol resistant) x viable recipient (chloramphenicol resistant) cfu.
The results clearly show that the gentamycin-resistance marker was acquired by P. putida KT2440 by both electrotransformation and conjugation, suggesting that the transposon was transferred to this non-enterobacterial recipient. In the electroporation experiment, the efficiency was low, with only 70 potential insertions per μg plasmid DNA, despite the fact that the strain had become very competent as shown by the high transformation efficiency achieved with a replicative plasmid. On the other hand, transposition frequency in the mating experiment was quite high, suggesting that conjugation may be a more efficient means to transfer the miniTn7BB-Gm transposon to P. putida. Similar results were obtained with the miniTn7BB-Gm transposon borne in the commercial plasmid pMA (Mr. Gene). Interestingly, conjugative transfer of pUC18 or pMA has not been described, and a transfer origin is not documented for any of these vectors. Similar frequencies and efficiencies have been obtained with other miniTn7 delivery plasmids, such as those of the pBK-miniTn7 series ([http://www.sciencedirect.com/science/article/pii/S0167701201002469 Koch et al., 2001]) (Fernando Govantes, personal communication).
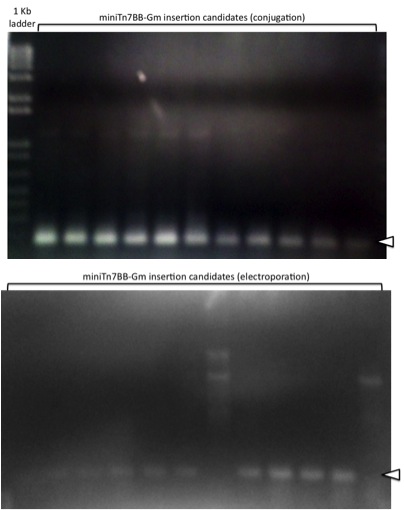
Site-specificity of the miniTn7-Gm insertions in P. putida KT2440 obtained was determined by PCR amplification using a primer annealing at the 3' end of glmS and a primer annealing at the Tn7R end. The occurence of a 164 bp product indicates successful site-specific integration at attTn7, while absence of this product suggests non-specific insertion elsewhere. 12 candidates each from the electroporation and mating experiments were tested by colony PCR as indicated (Figure 3). All candidates from the mating experiment and 10 out of 12 from the electroporation experiment displayed a band of the expected size, indicating that miniTn7BB-Gm efficiently inserts at the chromosomal attTn7 site of P. putida.
Figure 3. Agarose gels showing amplification from the chromosomal copies of miniTn7BB-Gm in candidates obtained in mating (top) or electroporation (bottom) experiments. Open arrowheads indicate the location of the correct 164 bp PCR products.
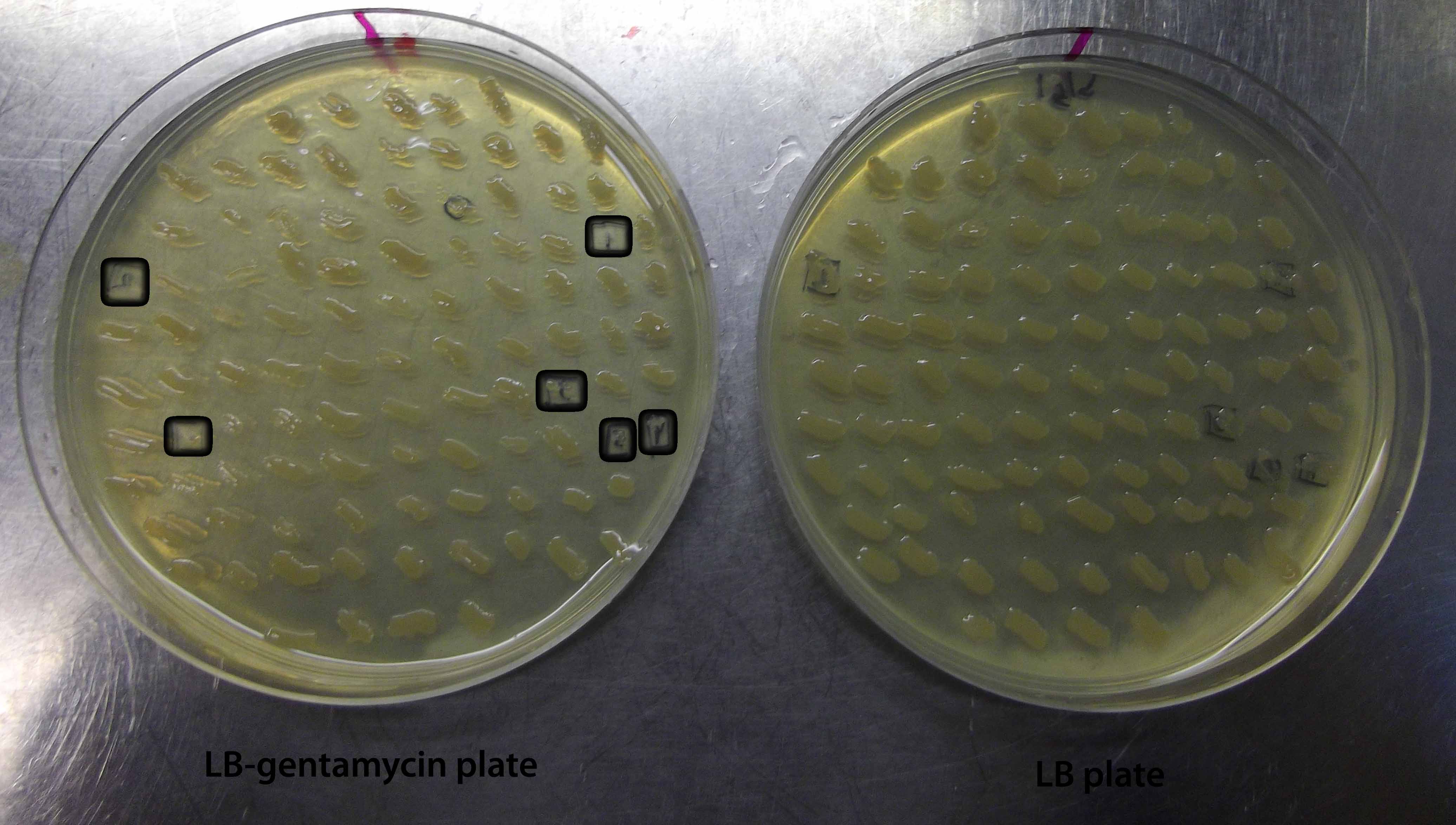
Finally, we tested whether the gentamycin resistance cassette in the miniTn7BB-Gm transposon can be excised by site-specific recombination performed by the flipase enzyme at the flanking FRT sites. To achieve this, we transferred the pFLP2 plasmid, encoding the yeast flipase, by electroporation into one of the P. putida clones bearing a miniTn7BB-Gm transposon insertion. Upon selection of the transformants on LB-carbenicillin plates, clones were patched onto LB-gentamycin and LB plates. 6 out of 100 candidates failed to grow on the LB-gentamycin plates, indicating the successful loss of the Gm-resistance cassette in the transposon. The low efficiency of excission may be improved by prolonged incubation on medium not containing gentamycin. Alternetively, the introduction of point mutations at the FRT elements to remove the XbaI restriction site may have resultee in decreased recognition of the target by the site-specific recombinase. Nevertheless, our results show that excission occurs and can be detected in a simple, effortless fashion (Figure 4).
Figure 4. Plates showing excision of the gentamycin resistance marker in P. putida KT2440 bearing miniTn7BB-Gm. Gentamycin-sensitive patches are boxed on both LB+gentamycin (left) and LB (right) plates.
Transposition of miniTn7BB-Gm in E. coli
In order to demonstrate miniTn7BB-Gm transposition into the E. coli DH5α chromosome, pUC18R6KT-miniTn7BB-Gm was transferred to this host by co-transformation with the helper plasmid pTNS2 of chemically competent cells [pTNS2 is not replicative in DH5α, as it harbors a R6K replication origin, but transiently produces the Tn7 transposition machinery ([http://http://www.nature.com/nmeth/journal/v2/n6/abs/nmeth765.html Choi et al. (2005)]))]. E. coli clones bearing miniTn7BB-Gm insertions were selected on LB plates supplemented with Gm. Viable counts were also performed, and plasmid transformation efficiency was determined using the gentamycin resistant replicative plasmid pBBR1mcs-5. The results are summarized in Table 2.
Table 2. Characterization of miniTn7BB-Gm transposition in E. coli DH5α. (a) Transformant (gentamycin resistant) cfu x μg-1 pBBR1mcs5. bcfu bearing a miniTn7BB-Gm insertion (gentamycin resistant) x μg-1 pUC18R6KT-miniTn7BB-Gm.
The results indicate that the gentamycin-resistant marker was acquired by the transformed E. coli DH5α cells, strongly suggesting that the transposon can be tranferred to an enterobacterial host by means of transformation of chemically competent cells. The efficiency with which transposon insertion candidates was obtained was somewhat (6-fold) higher than that obtained with P. putida, despite the fact that competence was clearly (6-fold) lower in the E. coli strain. This suggests that transformation is likely a better suited method for transposon delivery in E. coli than in P. putida.
Site-specificity of the miniTn7-Gm insertions obtained in E. coli DH5α was determined by PCR amplification as described above for P. putida. 11 out of 12 candidates tested in this fashion displayed the expected 164 bp band (data not shown), indicating that miniTn7BB-Gm efficiently inserts at the chromosomal attTn7 site of E. coli.
Conclusions
Taken together, the characterization of the miniTn7BB-Gm minitransposon indicates that this element is able to transpose efficiently in both an enterobacterial (E. coli), and a non-enterobacterial (P. putida) host, consistent with the previously described wide host-range of Tn7. The transposon can be delivered by means of electrotransformation, heat-shock transformation and conjugal mating from three different delivery vectors. We have also shown that most transposition events in both hosts occur in a site-specific fashion, at the known attTn7 sites present in their chromosomes. Finally, we have shown that the gentamycin resistance can be excised by flipase-dependent site-specific recombination at least in one of the hosts (P. putida), leaving an unmarked strain that may be suitable for applications in which the use of drug resistance markers is not acceptable.
Applications of BBa_K510000
Stable integration at a single site in the chromosome. Tn7 has the ability to integrate at a single target in the bacterial chromosome downstream from the conserved gene glmS. The insertion site is neutral, as it is located at a convergent intergenic region ([http://www.ncbi.nlm.nih.gov/pmc/articles/PMC211210/ Gringauz, 1988]; [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC287367/ Waddell, 1989]). Transposition is exerted by Tn7 transposase provided transiently from a suicide plasmid, granting a single transposition event and the impossibility of secondary transposition ([http://www.nature.com/nrm/journal/v2/n11/full/nrm1101-806a.html Peters, 2001]). Stable transposon insertion generates safer organisms, as horizontal gene transfer is minimized in this way.
Antibiotic selection not required. MiniTn7 insertions do not require antibiotic selection, as the transposon is faithfully replicated as part of the bacterial chromosome. Furthermore, all our miniTn7 vectors will be equipped with FRT-flanked antibiotic resistance that allows precise excision of the markers by Flp-mediated recombination ([http://www.ncbi.nlm.nih.gov/pubmed/7789817 Cherepanov, 1995]).
Suitability to multiple bacterial hosts. Most likely, Tn7 transposition is functional in a large array of Gram-negative bacteria. So far, Tn7 has been shown to transpose in over 20 different Gram-negative bacterial species, including several enterics, members of the genus Pseudomonas, Caulobacter crescentus, Desulfovibrio desulfuricans and others ([http://www.ncbi.nlm.nih.gov/pubmed/8556868 Craig, 1996]). Delivery of the transposon by mating or electroporation is possible in all of these organisms.
User Reviews
UNIQ11c8672267383a26-partinfo-00000000-QINU UNIQ11c8672267383a26-partinfo-00000001-QINU